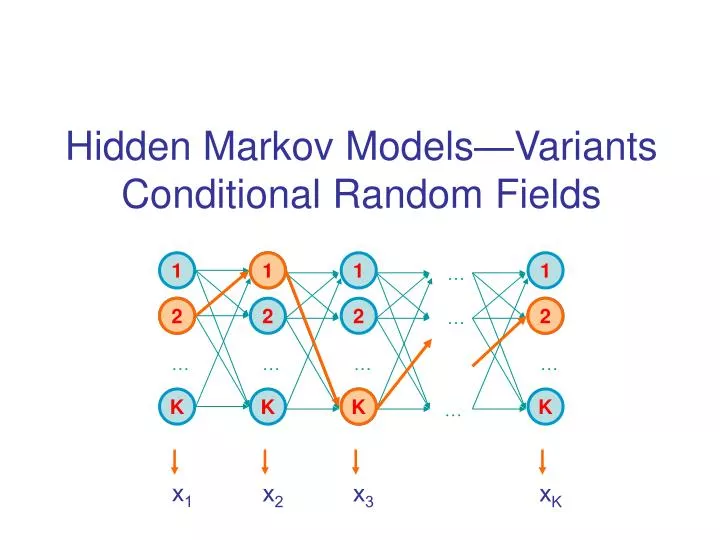
For example, let us consider the speech recognition problem, for which HMMs have been extensively used for several decades. Modeling observations in these two layers, one visible and the other invisible, is very useful, since many real world problems deal with classifying raw observations into a number of categories, or class labels, that are more meaningful to us. For this reason, an HMM is also called a doubly-embedded stochastic process. The hidden states form a Markov chain, and the probability distribution of the observed symbol depends on the underlying state. An HMM consists of two stochastic processes, namely, an invisible process of hidden states and a visible process of observable symbols. We call the observed event a `symbol' and the invisible factor underlying the observation a `state'. We show how these models and other types of HMMs can be employed in RNA sequence analysis.Ī hidden Markov model (HMM) is a statistical model that can be used to describe the evolution of observable events that depend on internal factors, which are not directly observable. Section 5 reviews context-sensitive HMMs (csHMMs) and profile context-sensitive HMMs (profile-csHMMs), which are especially useful for representing RNA families. 4, we focus on pair-HMMs and their applications in pairwise alignment, multiple sequence alignment, and gene prediction.
HIDDEN MARKOV MODEL MATLAB CODE EXAMPLE SOFTWARE
We also introduce publicly available profile-HMM software packages and libraries of pre-built profile-HMMs for known sequence families. Section 3 provides an overview of profile hidden Markov models and their applications. After reviewing the basic concept of HMMs, we introduce three types of HMM variants, namely, profile-HMMs, pair-HMMs, and context-sensitive HMMs, that have been useful in various sequence analysis problems.
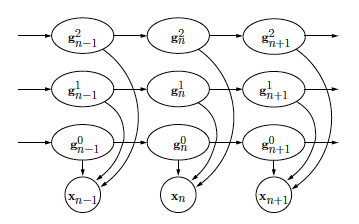
Algorithms for solving these problems are also introduced. 2, we begin with a brief review of HMMs and the basic problems that must be addressed to use HMMs in practical applications. The organization of the paper is as follows. In this paper, we give a tutorial review of HMMs and their applications in biological sequence analysis. For example, HMMs and their variants have been used in gene prediction, pairwise and multiple sequence alignment, base-calling, modeling DNA sequencing errors, protein secondary structure prediction, ncRNA identification, RNA structural alignment, acceleration of RNA folding and alignment, fast noncoding RNA annotation, and many others. Considering the remarkable success of HMMs in engineering, it is no surprise that a wide range of problems in biological sequence analysis have also benefited from them. HMMs are well-known for their effectiveness in modeling the correlations between adjacent symbols, domains, or events, and they have been extensively used in various fields, especially in speech recognition and digital communication. Until now, various signal processing models and algorithms have been used in biological sequence analysis, among which the hidden Markov models (HMMs) have been especially popular. Given the expanding list of newly sequenced genomes and the increasing demand for genome re-sequencing in various comparative genomics projects, the importance of computational tools in biological sequence analysis is expected to grow only further. In order to extract meaningful information from the data, we need computational techniques that can efficiently analyze the data according to sound mathematical principles. However, considering the huge size of the available data, it is virtually impossible to analyze them without the help of computational methods.

The sequenced genomes contain a wealth of invaluable information that can help us better understand the underlying mechanisms of various biological functions in cells. The successful completion of many genome sequencing projects has left us with an enormous amount of sequence data.
